Access to software and other resources can be utilized by clicking the corresponding links below
ASDP
ASDP is an automated NMR NOE as signment engine. It uses a distinct bottom-up topology-constrained approach for iterative NOE interpretation and generates 3D structures of the protein that is as close to the true structure as possible.
To cite ASDP
1. Huang, Y.J., Tejero, R., Powers, R. and Montelione, G.T. A topology-constrained distance network algorithm for protein structure determination from NOESY data. PROTEINS: Structure, Function, Bioinformatics, 62: 587-603, 2006.
2. Huang, Y.J., Mao, B., Xu, F., and Montelione, G.T. Guiding automated NMR structure determination using a global optimization metric, the NMR DP score. J. Biomol. NMR, 62: 439-451, 2015.
AutoAssign
AutoAssign is a constraint-based expert system for automating the analysis of backbone resonance assignments using NMR spectra.
To cite AutoAssign
1. Zimmerman, D.E., Kulikowski, C.A., Huang, Y., Feng, W., Tashiro, M., Shimotakahara, S., Chien, C., Powers, R., and Montelione, G.T. Automated analysis of protein NMR assignments using methods from artificial intelligence. J. Mol. Biol, 269: 592-610, 1997.
2. Moseley, H.N., Monleon, D., Montelione, G.T. Chapter 6 Automatic determination of protein backbone resonance assignments from triple resonance nuclear magnetic resonance data. Methods in Enzymology, Vol. 339 Eds. T.J. James, V. Dotsch and U. Schmitz. Academic Press. San Diego, CA, 2001

Dismeta- Disorder prediction meta-server
Dismeta polls a number of disorder predictors and reports the results of each. Consensus disorder is plotted per residue.
To cite DisMeta
1. Huang, Y.J., Acton, T.B., Montelione, G.T. Chapter 1. DisMeta: A meta server for construct design and optimization. Methods in Molecular Biology, Structural Biology, Vol. 1091: Ed. JM Walker, Humana Press. New York, NY, 2014.
2. Acton, T.B., Xiao, R., Anderson, S., Aramini, J., Buchwald, W.A., Ciccosanti, C., Conover, K., Everett, J., Hamilton, K., Huang, Y.J., Janjua, H., Kornhaber, G., Lau, J., Lee, D.Y., Liu, G., Maglaqui, M., Ma, L., Mao, L., Patel, D., Rossi, P., Sahdev, S., Shastry, R., Swapna, G.V., Tang, Y., Tong, S., Wang, D., Wang, H., Zhao, L. and Montelione, G.T. Chapter 2 Preparation of protein samples for NMR structure, function, and small-molecule screening studies. Methods in Enzymology, Vol. 493: Ed. LC Kuo. Academic Press. San Diego, CA, 2011.

NESG NMR wiki
The NESG wiki shares experimental protocols as well as training and educational materials in the fields of structural biology, structural genomics and biomolecular NMR.
NMR 2.0
NMR 2.0 provides a collection of collaborative and instructive tools to advance NMR studies.

PDBStat
Provides extensive tools for assessing NMR structures against data, including restraint violation analysis and RDC analysis, as well as tools for converting between coordinate and restraint formats, including NEF format, structural superimposition, and contact map generation.
To cite PDBStat
1. Tejero, R., Snyder, D., Mao, B., Aramini, J.M., and Montelione, G.T. PDBStat: A universal restraint converter and restraint analysis software package for protein NMR. J. Biomol. NMR, 56: 337-351, 2013.
PSVS (Protein Structure Validation Software Suite)
PSVS is used for assessment of protein structures generated from NMR, X-ray crystallographic and homology modeling methods.
To cite PSVS
1. Bhattacharya, A., Tejero, R. and Montelione, G.T. Evaluating protein structures determined by structural genomics consortia. PROTEINS: Structure, Function, Bioinformatics, 66: 778-795, 2007
RPF (Recall, Precision, and F-measure scores)
Structure quality assessment measurements based on information retrieval statistics
To cite RPF
1. Huang, Y.J.; Powers, R.; Montelione, G.T. J. Am. Chem. Soc. 2005 127: 1665-1674. Protein NMR recall, precision, and F-measure scores (RPF scores): Structure quality assessment measures based on information retrieval statistics.
2. Huang, Y.J., Rosato, A., Singh, G., and Montelione, G.T. RPF: a quality assessment tool for protein NMR structures. Nucleic Acids Research, 40: W542-546, 2012.
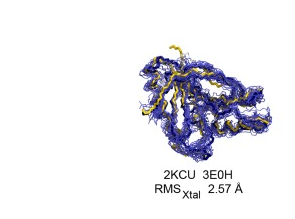
NMR / X-ray Structure Pair Data Repository
NMR spectroscopy and X-ray diffraction data for 41 “NMR / X-ray” structure pairs determined using conventional triple-resonance NMR methods with extensive sidechain resonance assignments . In addition, several NMR data sets for perdeuterated, methyl-protonated protein samples are included in this repository. The NMR / X-ray Structure Pair Data Repository provides a valuable resource for new computational NMR methods development.
A community resource of experimental data for NMR / X-ray crystal structure pairs
Primer Prim’er
Primer Prim’er designs PCR primer sets for endonuclease
and viral recombinase based cloning strategies.
pXs (probability of crystal structure) calculator
pXs calculates the probability of a given protein sequence to yield a X-Ray structure.

Structure Superposition with FindCore and PDBstat
A structure superimposition server using ordered residues or core residues, based on PdbStat and FindCore respectively.
1. Snyder, D.A. and Montelione, G.T. Clustering algorithms for identifying core atom sets and for assessing the precision of protein structure ensembles. PROTEINS: Structure, Function, Bioinformatics, 59: 673-686, 2005.
2. Snyder, D.A., Grullon, J., Huang, Y.J., Tejero, R. and Montelione, G.T. The expanded FindCore method for identification of a core atom set for assessment of protein structure prediction. PROTEINS: Structure, Function, Bioinformatics, 82 Suppl 2: 219-230, 2014.
AVS- Chemical Shift Assignment Validation Server
AVS is used to validate chemical shift data, flagging shifts that are outside the range typically observed in proteins.
To cite Assignment Validation Server
1. Moseley, H.N., Sahota, G. and Montelione, G.T. Assignment validation software suite for the evaluation and presentation of protein resonance assignment data. J. Biomol. NMR, 28: 341-355, 2004